In certain instances, “open sesame” might be something you exclaim to magically open the door to a cave full of treasure, but for the sesame plant, open sesame is a way of life. In sesame’s case, seeds are the treasure, which are kept inside a four-chambered capsule. In order for the next generation of plants to have a chance at life, the seeds must be set free. Sesame’s story is similar to the stories of numerous other plant species whose seeds are born in dehiscent fruits. But in this instance, the process of opening those fruits is fairly unique.
Sesamum indicum is a domesticated plant with a 5000 plus year history of cultivation. It shares a genus with about 20 other species – most of which occur in sub-Saharan Africa – and belongs to the family Pedaliaceae – the sesame family. Sesame was first domesticated in India and is now grown in many other parts of the world. It is an annual plant that is drought and heat-tolerant and can be grown in poor soils and locations where many other crops might struggle. However, the best yields are achieved on farms with fertile soils and adequate moisture.

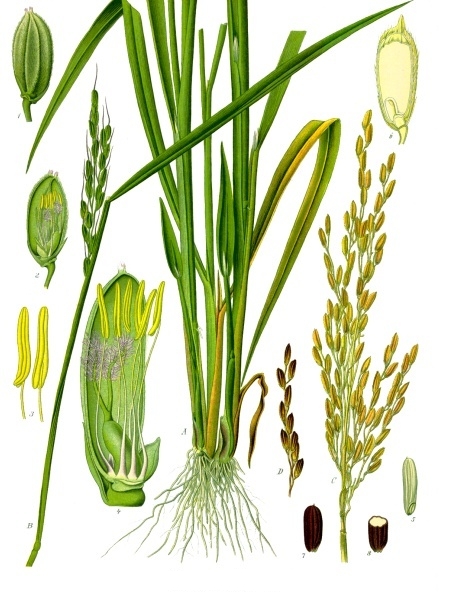
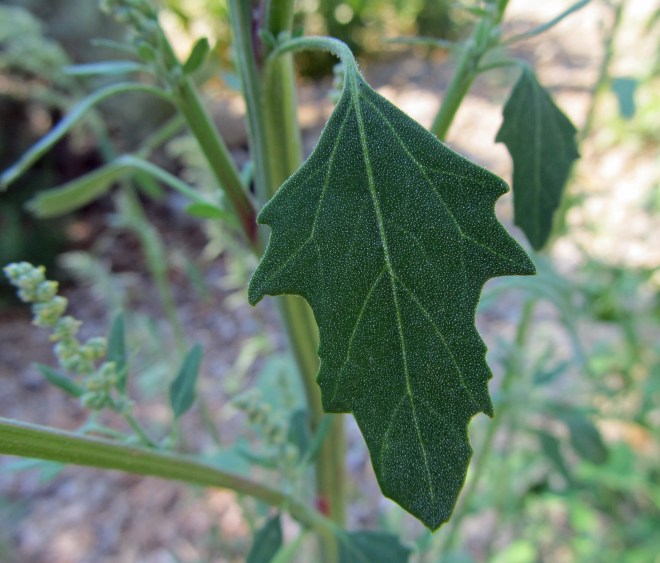
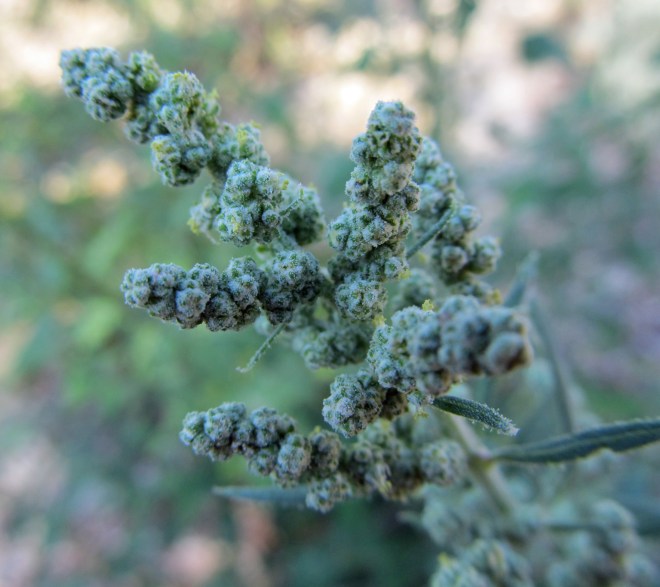
Depending on the variety and growing conditions, sesame can reach up to 5 feet tall and can be unbranched or highly branched. Its broad lance-shaped leaves are generally arranged directly across from each other on the stem. The flowers are tubular, similar in appearance to foxglove, and are typically self-pollinated and short-lived. They come in shades of white, pink, blue, and purple and continue to open throughout the growing season as the plant grows taller, even as fruits formed earlier mature. The fruits are deeply-grooved capsules with at least four separate chambers called locules. Rows of tiny, flat, teardrop shaped seeds are produced in each chamber. The seeds are prized for their high oil content and are also used in numerous other ways, both processed and fresh. One of my favorite uses for sesame seeds is tahini, which is one of the main ingredients in hummus.
The fruits of sesame are dehiscent, which means they naturally split open upon reaching maturity. Compare this to indehiscent fruits like acorns, which must either rot or be chewed open by an animal in order to free the seeds. Dehiscence is also called shattering, and in many domesticated crop plants, shattering is something that humans have selected against. If fruits dehisce before they can be harvested, seeds fall to the ground and are lost. Selecting varieties that hold on to their seed long enough to be harvested was imperative for crops like beans, peas, and grains. In domesticated sesame, the shattering trait persists and yield losses are often high.

Most of the world’s sesame crop is harvested by hand. The plants are cut, tied into bundles, and left to dry. Once dry, they are held upside down and beaten in order to collect the seeds from their dehisced capsules. When harvested this way, naturally shattering capsules may be preferred. But in places like the United States and Australia, where mechanical harvesting is desired, it has been necessary to develop new, indehiscent varieties that can be harvested using a combine without losing all the seed in the process. Developing varieties with shatter-resistant seed pods, has been challenging. In early trials, seed pods were too tough and passed through threshers without opening. Additional threshing damaged the seeds and caused the harvest to go rancid. Mechanically harvested varieties of sesame exist today, and improvements in these non-shattering varieties continue to be made.
In order to develop these new varieties, breeders have had to gain an understanding of the mechanisms behind dehiscence and the genes involved in this process. This research has helped us appreciate the unique way that the capsules of the sesame plant dehisce. As in the seed bearing parts of many other plant species, the capsules of sesame exhibit hygroscopic movements. That is, their movements are driven by changes in humidity. The simplest form of hygroscopic movement is bending, which can be seen in the opening and closing of pine cone scales. A more complex movement can be seen in the seed pods of many species in the pea family, which both bend and twist as they split open. In both of these examples, water is evaporating from the plant part in question. As it dries it bends and/or twists, thereby releasing its contents.

The cylindrical nature and cellular composition of sesame fruits leads to an even more complex form of hygroscopic movement. Initially, the capsule splits at the top, creating an opening to each of the four locules. The walls of each locule bend outward, then split and twist as the seed falls from the capsule. In a study published in Frontiers in Plant Science (2016), researchers found that differences in the capsule’s inner endocarp layer and outer mesocarp layer are what help lead to this interesting movement. The endocarp layer is composed of both transvere (i.e. circumferential) and longitudinal fiber cells, while the mesocarp is made up of soft parenchyma cells. The thicknesses of these two layers gradually changes along the length of the capsule. As the mesocarp dries, the capsule initially splits open and starts bending outwards, but as it does, resistance from the fiber cells in the endocarp layer causes further bending and twisting (see Figure 1 in the report for an illustration). As the researchers write, “the non-uniform relative thickness of the layers promotes a graded bi-axial bending, leading to the complex capsule opening movement.”
All this considered, a rock rolling away from the entrance of a cave after giving the command, “Open sesame!” almost seems simpler than the “open sesame” experienced by the fruit of the sesame plant.
See Also: Seed Shattering Lost – The Story of Foxtail Millet




![Grapefruit (Citrus x paradisi) - A hybrid between sweet orange (Citrus sinensis) and shaddock (Citrus maxima) that "occurred far beyond the region of domestication and rather recently [the 18th centruy]." (photo credit: wikimedia commons)](https://awkwardbotany.com/wp-content/uploads/2014/12/grapefruit.jpeg?w=660)